Content:
Diffraction limits & principal axes of ellipsoid fitted to diffraction cut-off surface:
1.911 0.9497 0.0000 0.3131 0.999 a* + 0.034 c*
1.916 0.0000 1.0000 0.0000 b*
1.869 -0.3131 0.0000 0.9497 -0.539 a* + 0.843 c*
| Date deposited | Date data collection | Resolution | R, Rfree |
| 20060714 | 20060603 | 1.95 | 0.2020 0.2210 |
Molprobity (CCP4 7.0 version) summary:
Ramachandran outliers = 0.00 %
favored = 98.36 %
Rotamer outliers = 1.13 %
C-beta deviations = 0
Clashscore = 6.48
RMS(bonds) = 0.0152
RMS(angles) = 1.81
MolProbity score = 1.40
Resolution = 1.95
R-work = 0.2020
R-free = 0.2210
Additional analysis:
Number of waters = 308 <B> (all atoms) = 39.93 ( sd = 9.12 ) for 2728 non-hydrogen atoms <B> (protein) = 38.35 ( sd = 7.64 ) for 2391 non-hydrogen atoms <B> (water) = 49.54 ( sd = 9.43 ) for 308 non-hydrogen atoms <B> (others) = 68.48 ( sd = 9.20 ) for 29 non-hydrogen atoms B min/max (all non-hydrogen atoms) = 23.27 / 82.17 B min/max (protein non-hydrogen atoms) = 23.27 / 76.95 B min/max (water non-hydrogen atoms) = 28.15 / 77.13 B min/max (other non-hydrogen atoms) = 54.66 / 82.17
Molprobity (CCP4 7.0 version) summary:
Ramachandran outliers = 0.00 %
favored = 98.69 %
Rotamer outliers = 1.51 %
C-beta deviations = 0
Clashscore = 3.13
RMS(bonds) = 0.0118
RMS(angles) = 1.55
MolProbity score = 1.24
Resolution = 1.95
R-work = 0.1883
R-free = 0.2143
Additional analysis:
Number of waters = 320 <B> (all atoms) = 40.31 ( sd = 9.91 ) for 2740 non-hydrogen atoms <B> (protein) = 38.63 ( sd = 8.45 ) for 2391 non-hydrogen atoms <B> (water) = 51.87 ( sd = 11.75 ) for 320 non-hydrogen atoms <B> (others) = 51.46 ( sd = 6.54 ) for 29 non-hydrogen atoms B min/max (all non-hydrogen atoms) = 23.45 / 93.65 B min/max (protein non-hydrogen atoms) = 23.45 / 83.79 B min/max (water non-hydrogen atoms) = 27.65 / 93.65 B min/max (other non-hydrogen atoms) = 41.64 / 61.04
Refinement progression:
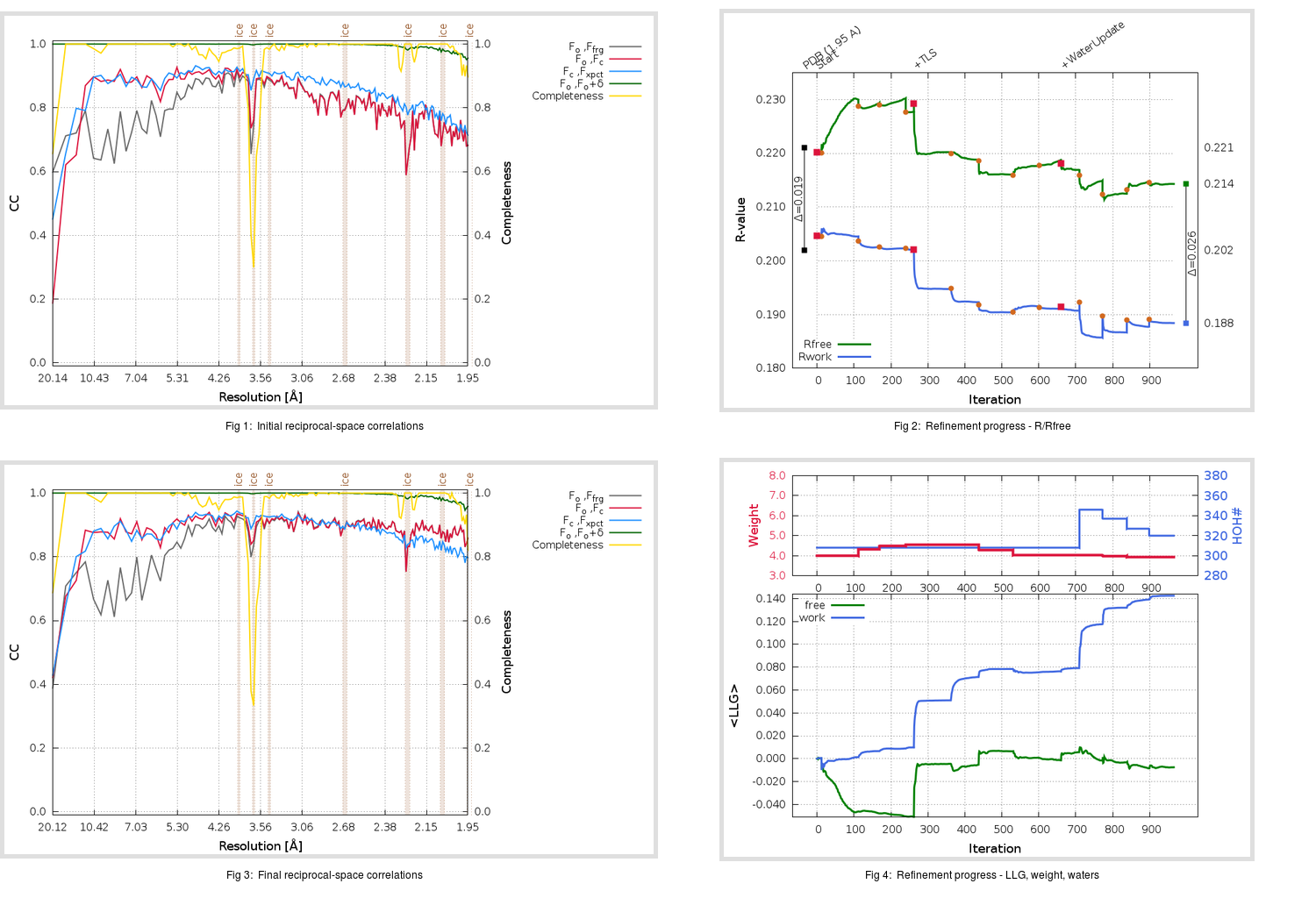 |
Results:
| File | Remark |
| 2HOB_aB_refine.01_03_refine.pdb.gz | exact refinement commands are in header |
| 2HOB_aB_refine.01_03_refine.mtz.gz | including original deposited data and several re-refinement map coefficients |
| 2HOB_aB_refine.01_03_BUSTER_model.cif.gz | including any non-standard compound restraints |
| 2HOB_aB_refine.01_03_BUSTER_refln.cif.gz |